In recent years investigators have successfully developed super-efficient systems using alternative technologies to generate genetically engineered mice and rats much faster and cheaper* compared to traditional targeted mutations methods: Zinc Finger Nucleases (ZFNs), Transcription Activator-Like Effector Nucleases (TALENs) and CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) /Cas (CRISPR associated) systems. For rats, in the absence of efficient germ line competent ES cells, nucleases were the only reproducible way to target genes.
Nucleases are powerful tools to edit and modify the genome. The different reported systems allow to create specific double-strand breaks at the target locus that trigger DNA repair mechanisms. These corrections result in i) knock-outs through Non-Homologous End Joining (NHEJ) and ii) knock-in through Homologous Directed Repair. The NHEJ pathway is the favored correction mechanism. The last developed genome editing technology is the CRISPR/Cas9 technology (Our Guidelines:
 FAQs CRISPR (868.6 KB)
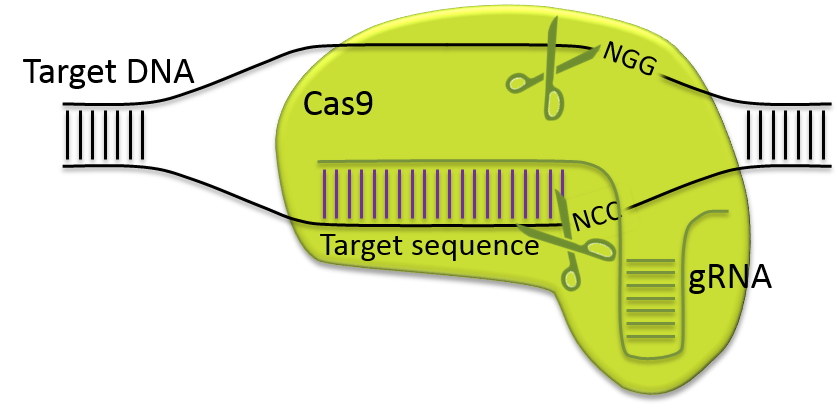
). This system, inherited from a prokaryotic defense mechanism, consists of a Cas endonuclease that is directed to cleave a target sequence by a guide RNA (gRNA). Using the CRISPR/Cas9 nuclease system, modifications can be introduced by injecting directly Cas9 mRNA or protein and gRNA into fertilized oocytes.
FAQs CRISPR (868.6 KB)
). This system, inherited from a prokaryotic defense mechanism, consists of a Cas endonuclease that is directed to cleave a target sequence by a guide RNA (gRNA). Using the CRISPR/Cas9 nuclease system, modifications can be introduced by injecting directly Cas9 mRNA or protein and gRNA into fertilized oocytes.
PHENOMIN has implemented electroporation for the delivery of CRISPR/Cas9 with or without small donor DNA. This method allows the drastic reduction of the number of donor females and fertilized oocytes. We consistently obtained high efficiency mouse genome editing for multiple genes, and we successfully generated mouse embryos with a variety of editing schemes, including indel mutations, point mutations, genomic deletions, and small precise insertions. Taken together, CRISPR-electroporation is a simple, economic, 3Rs friendly, high throughput, and highly efficient technique for genome editing in vivo, which can replace the traditional microinjection-dependent genome editing technique in mice and in other mammalian species (
 Generation of mutated mice combining CRISPR/Cas9 and electroporation (480.9 KB)
).
Generation of mutated mice combining CRISPR/Cas9 and electroporation (480.9 KB)
).
* Not true yet for complex models but it will very likely evolve.
.
|
|
| |
|
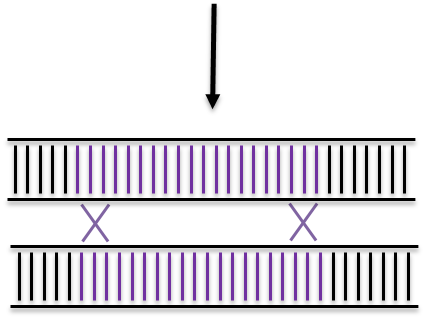
CRISPR/Cas9 targeted double strand break
|
|
|
|
|
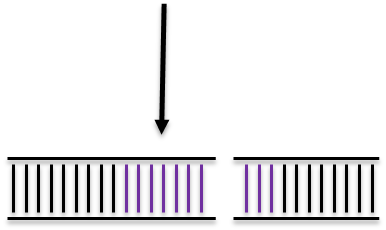
Non Homologous End Joining
|
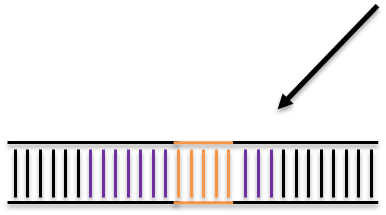
Homologous Recombination
|
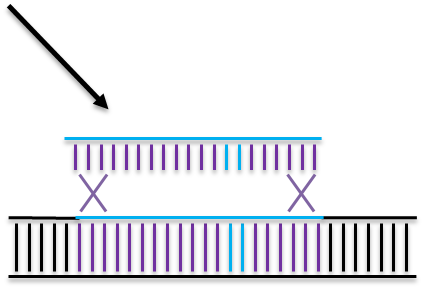
Homologous Directed Repair
|
|
Design, synthesis of targeting gRNA and Cas9
|
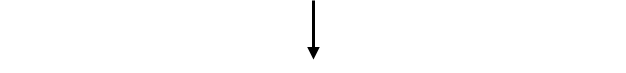
Injection into fertilized oocytes
|

|
Implantation into pseudo pregnant female
|
| |
|
|
|
F0 breeding with WT animal
|
Sequencing of all Founders and selection
|
Mosaic F0 animals
|

|
|
|
 Heterozygous F1 animals
Heterozygous F1 animals
|
Sequencing of all F1
|
New alleles can be found
Some alleles are not transmitted to F1 generation
|
Advantages
- Target design simplicity: gRNAs can be designed to target nearly any sequence in the genome; improved bioinformatics tools help to identify the most appropriate sequences to design gRNAs.
- Time-saving:
- Synthetizing a gRNA is simplier and faster than an homologous recombination vector for ES cells.
- Modifications can be introduced by material injection into fertilized eggs. This eliminates the long process of electroporation and selection of mouse ES cells required to create targeted mutant mice using classical homologous recombination techniques.
- High efficiency: in our hands, for knock-out models, the recombination frequency is boosted hundred(s)-fold(s) compared to the classical homologous recombination techniques. This advantage can be used to highly improve homologous recombination efficiency in ES cells (coelectroporation of standard plasmid RH with Cas9 and gRNA plasmid) and generate large replacement that were not (easily) feasible previously.
- Cost-effective:
- Time to germ line transmission is reduced compared to homologous recombination.
- The CRISPR/Cas system does not need to be paired with separate cleaving enzymes as other tools do. RNAs encoding both the Cas9 protein and gRNA(s) can be injected.
- Works in all mouse and rat genetic backgrounds. As in classical transgenesis experiments (random insertion of transgene by pronuclear injection), the critical point is the availability, robustness and capacity of development of embryos from inbred or outbred strains, as well as microinjectionist proficiency.
Drawbacks
- Off-target effects: Cas9 complexes with a sgRNA can cleave DNA sequences that contains base mismatch(es) with the intended target sequence, especially in the 5’ region. Therefore, mutations can be introduced at non-specific loci with similar, but not identical, homology to the target site. Different softwares - we use CRISPOR - allow now to search for the best sgRNA with the lowest off-target potential and reduce dramatically the risk.
- Mosaicism and multiple alleles: as the nucleases may not necessarily cut the DNA at the one cell stage of embryonic development, many different allele variants can be found in one single founder. Furthermore, the NHEJ process can produce different mutations. These events make elucidating the genotype of founders very challenging; the production of mice or rats with multiple variants also creates phenotyping bottlenecks. Therefore breeding is required to segregate and isolate animals that carry single mutations.
- Multiple site integration at the locus: when the donor DNA is a circular plasmid, the targeted alleles often carry multiple integrations of the targeting vector (Pavlovic et al., 2016); concatemers are also observed with linear DNA.
PHENOMIN can perform a deeper characterization of CRISPR-edited animals. Please, do not hesitate to contact us.
- Not yet suitable for complex genome modifications: CRISPR/Cas9 in embryos works extremely well for the generation of simple alleles (constitutive knock out, knock-in of point mutations), but is not the technology of choice for the introduction of more complex modifications. Although PHENOMIN and others have successfully introduced complex modifications in the murine genome using CRISPR/Cas9 in embryos the complexity of validation procedures for these projects can result in increased timelines and costs.
For the moment, we recommend using ES cells (with or without the help of CRISPR/Cas9) to generate complex genomic modifications or large knock-in. This approach allows a perfect validation of the targeted alleles at the ES stage and is in accordance with the principles of the 3Rs (Replacement, Reduction and Refinement).
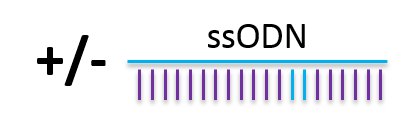
- Random single strand DNA integration: using droplet digital PCR, we were able to show that ssODN can integrate randomly in the genome and should be evaluated.
PHENOMIN expertise
- More than 100 mouse and rat models realized with the CRISPR/Cas9 technology.
- PHENOMIN has brought a large contribution to the establishment of CRISPR guidelines in the frame of CELPHEDIA network:
 FAQs CRISPR (868.6 KB)
FAQs CRISPR (868.6 KB)
- Our microinjectionists determine the best approach to inject the nuclease and the gRNA, depending on the specificities of your project and objectives. The Cas9 nuclease can be delivered as mRNA or protein; the single guides are delivered RNA. RNAs, proteins and DNA (oligonucleotides, plasmids or single stranded DNA) can be injected either into the cytoplasm or into the nucleus. PHENOMIN has implemented electroporation for the delivery of CRISPR/Cas9 with or without small donor DNA. This method allows the drastic reduction of the number of donor females and fertilized oocytes. CRISPR-electroporation is a simple, economic and 3Rs friendly technique for genome editing in vivo ( Generation of mutated mice combining CRISPR/Cas9 and electroporation ).
- PHENOMIN has evaluated the last Base Editing and conclude that standard CRISPR/Cas9 is still the most appropriate approach to obtain any kind of models.
- PHENOMIN can perform a deep chararcterization of CRISPR-edited animal to overcome the multiple site integration at the locus effect.
- PHENOMIN team’s expertise in CRISPR technology allowed the development of structural variant models using the innovative CRISMERE technology (large deletions, duplications and inversions) (Birling et al., 2017).
- PHENOMIN uses its experience to determine the most appropriate technology (nuclease-based or ES cell-based) for each project.
- Implication in international consortia (near 20 different consortia) to benefit from the latest technics and scientific expertise
- Continuous R&D activity to improve our technics and success rate and propose our customers the best approach for their projects.
- Personalized genome engineering service to fit our customer needs.
PHENOMIN experts webinars
PHENOMIN experts publications

![]() FAQs CRISPR (868.6 KB)
). This system, inherited from a prokaryotic defense mechanism, consists of a Cas endonuclease that is directed to cleave a target sequence by a guide RNA (gRNA). Using the CRISPR/Cas9 nuclease system, modifications can be introduced by injecting directly Cas9 mRNA or protein and gRNA into fertilized oocytes.
FAQs CRISPR (868.6 KB)
). This system, inherited from a prokaryotic defense mechanism, consists of a Cas endonuclease that is directed to cleave a target sequence by a guide RNA (gRNA). Using the CRISPR/Cas9 nuclease system, modifications can be introduced by injecting directly Cas9 mRNA or protein and gRNA into fertilized oocytes.![]() Generation of mutated mice combining CRISPR/Cas9 and electroporation (480.9 KB)
).
Generation of mutated mice combining CRISPR/Cas9 and electroporation (480.9 KB)
).












 Heterozygous F1 animals
Heterozygous F1 animals
